Computational Biophysics
Modeling Biomolecular Behavior to Power Next-Generation Biomedical Discoveries
Computational Biophysics: A Foundation for Modern Scientific Discovery
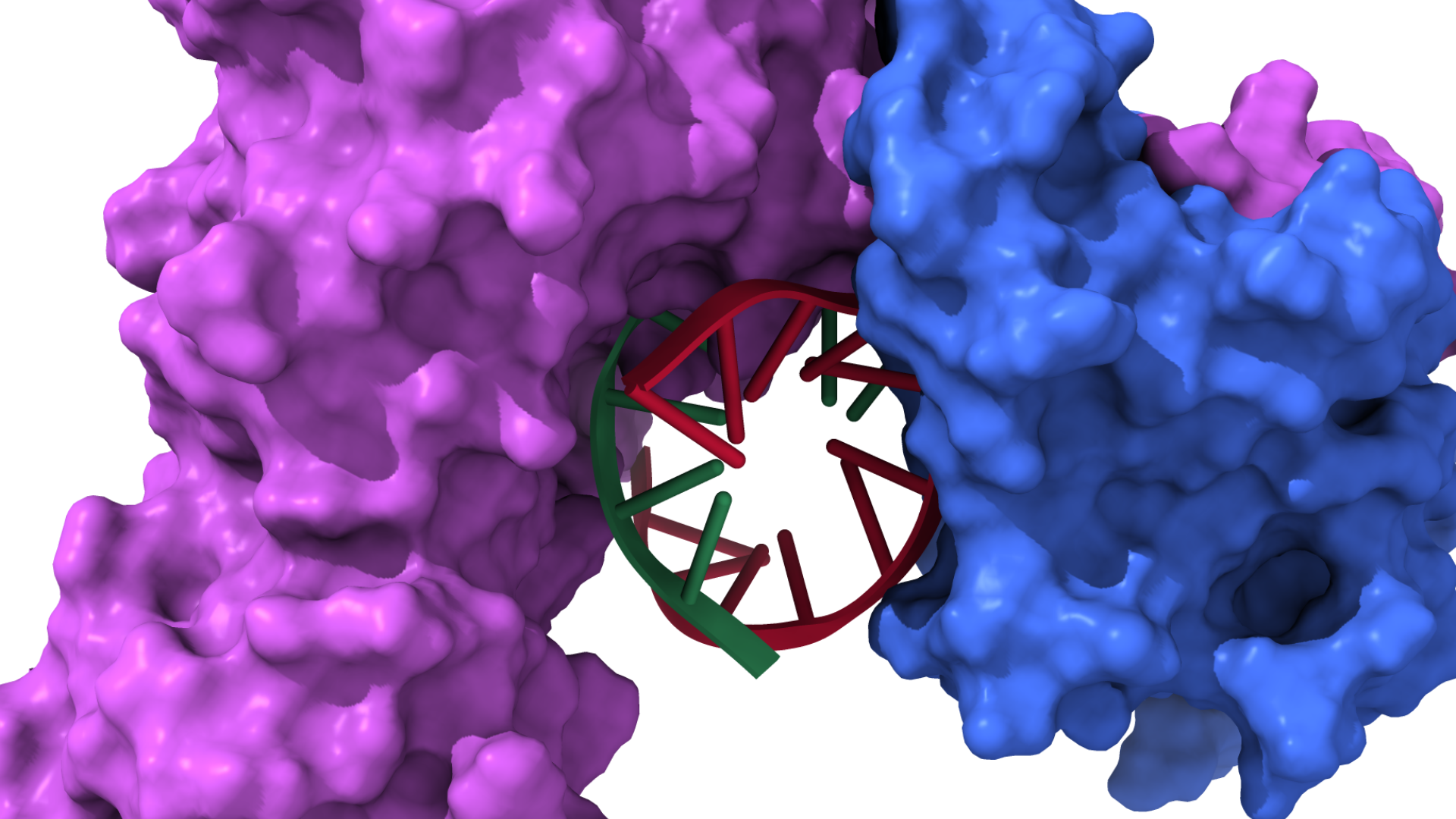

Computational Biophysics is an interdisciplinary field that applies advanced computational methods to investigate the physical principles governing biological systems. By combining physics, chemistry, biology, and computer science, the field enables researchers to study molecular structures, interactions, and dynamics with unprecedented detail. Computational models and simulations bridge the gap between theoretical understanding and experimental observation, providing insights into biomolecular processes that are often difficult or impossible to capture using laboratory techniques alone.
A central component of computational biophysics is Molecular Dynamics (MD) simulation, a technique that models the behavior of atoms and molecules over time. Using classical mechanics—most notably Newton’s laws of motion—MD simulations track how particles move and interact at the atomic scale. These simulations reveal fundamental information about structure, stability, conformational changes, and reaction pathways in biological and material systems. Because of their high-resolution nature, MD simulations have become indispensable in exploring complex biomolecular processes such as protein folding, membrane dynamics, enzyme mechanisms, and nucleic acid behavior.
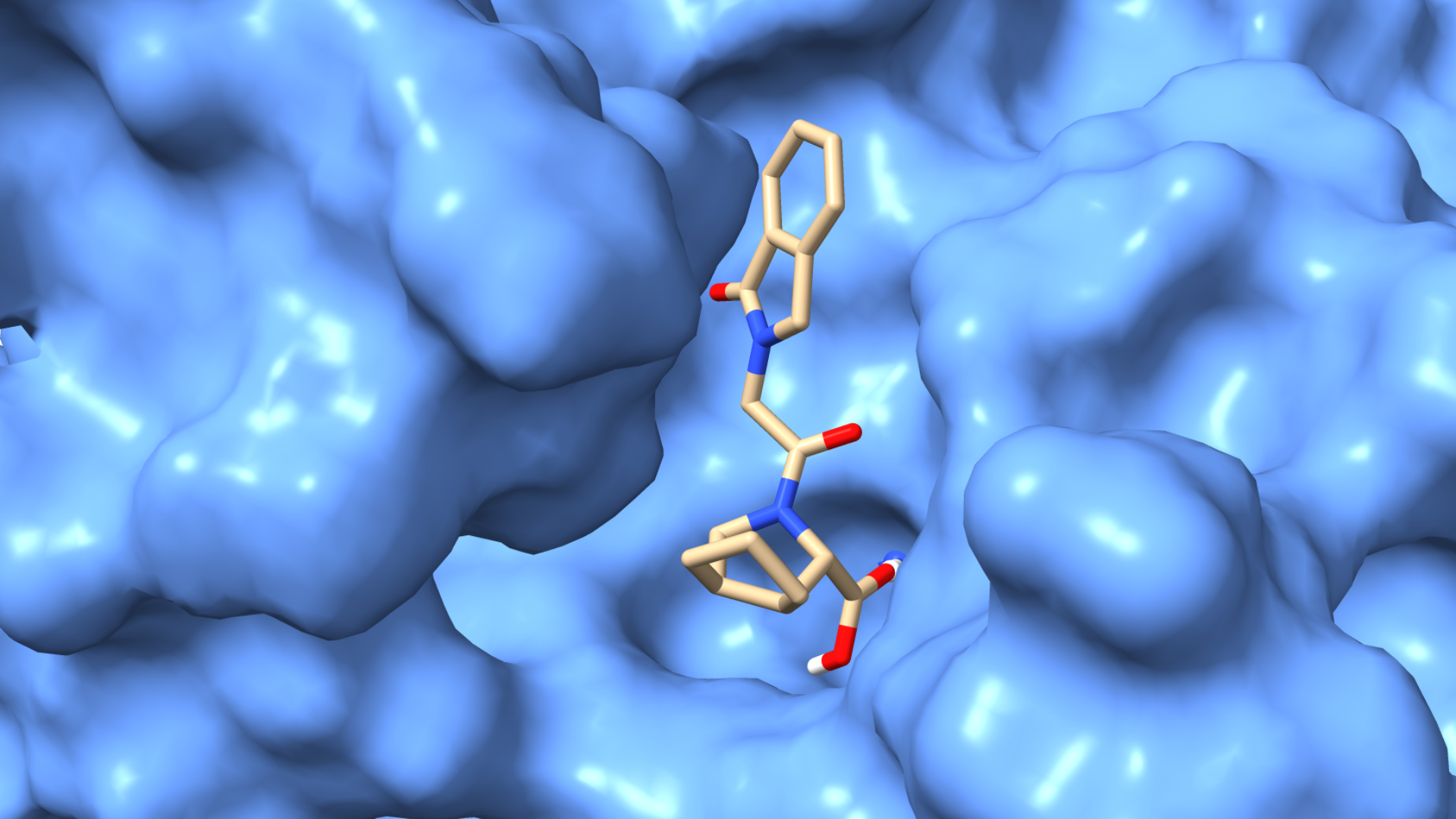
Another key area is virtual screening, a computational approach used to evaluate how small molecules, such as potential drugs, interact with target proteins. Virtual screening relies heavily on molecular docking to predict the strength and orientation of these interactions. By rapidly assessing thousands to millions of compounds in silico, it dramatically reduces the time, cost, and experimental workload associated with early-stage drug discovery. This technique not only identifies promising lead compounds but also supports drug repurposing and the development of personalized therapeutic strategies.
The Role of Computational Biophysics in Predicting Cancer Solutions
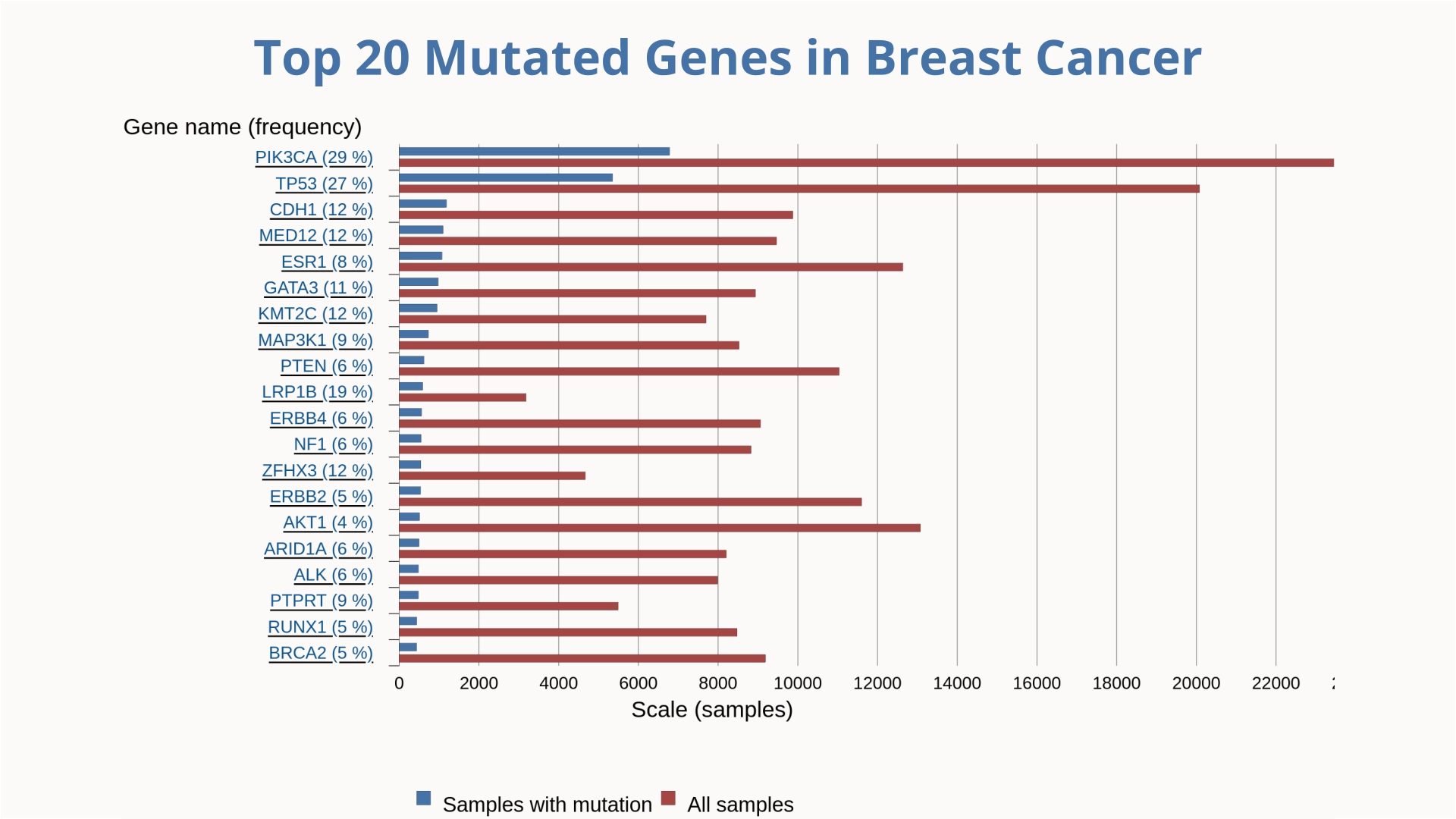
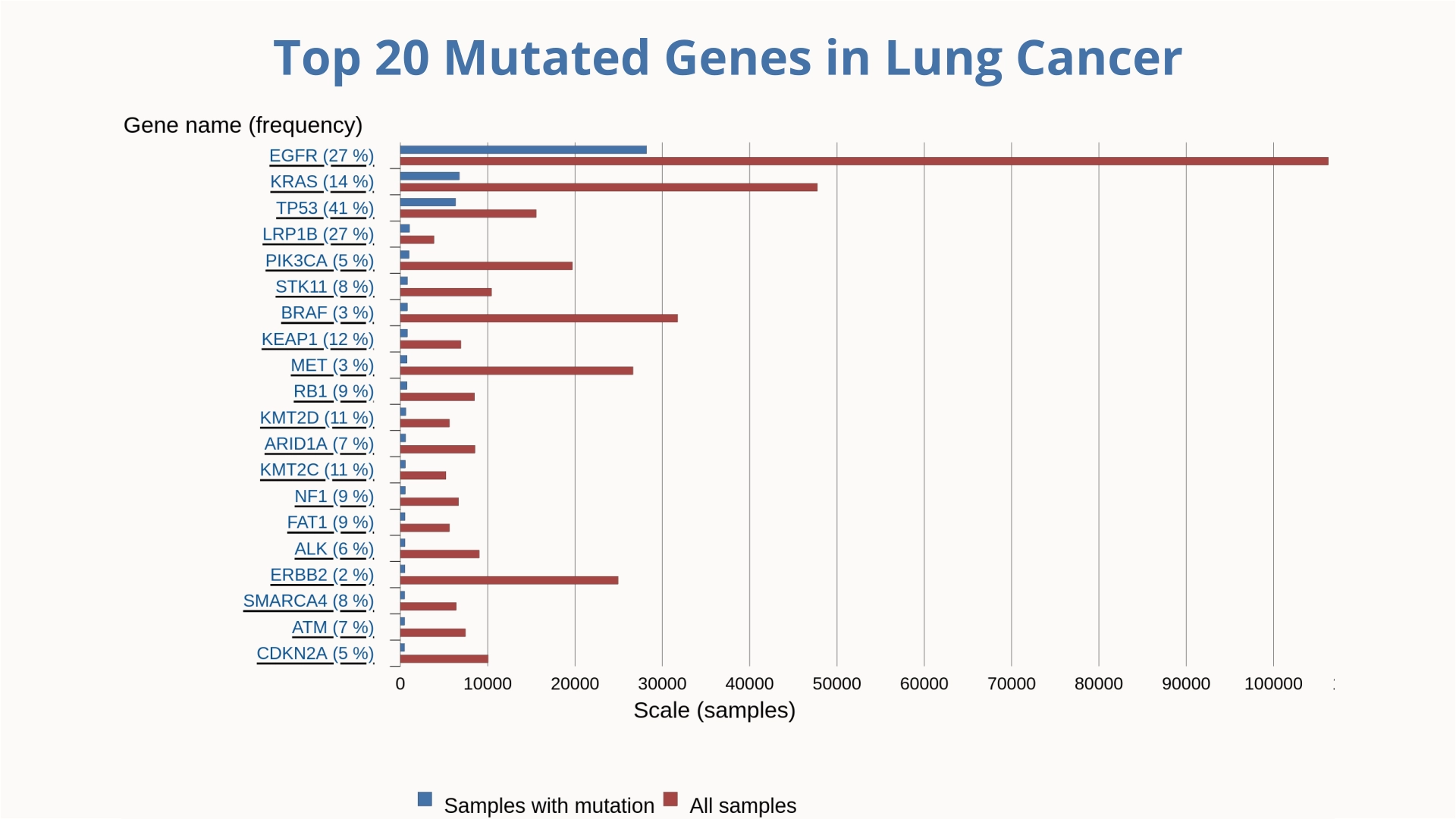
Lung cancer remains the leading cause of cancer-related deaths among men globally, while breast cancer holds the same position among women. Understanding the molecular biology of cancer requires identifying and analyzing the mutations that drive these diseases.
Understanding the genetic landscape of cancer is essential for identifying disease drivers and developing targeted therapies. The previous charts highlight the top 20 most frequently mutated genes in breast cancer and lung cancer, offering a comparative view of key genetic alterations across these two major cancer types. Visualizing mutation patterns helps researchers pinpoint critical pathways involved in tumor initiation and progression, ultimately guiding precision medicine and the design of more effective treatment strategies.
Computational biophysics plays a vital role in transforming these genomic insights into actionable solutions. By combining molecular dynamics simulations, virtual screening, and structure-based modeling, researchers can investigate how cancer-associated mutations alter protein function at the atomic level. These tools enable the prediction of vulnerable molecular sites, the identification of potential drug candidates, and the design of tailored therapeutic strategies that specifically target mutated proteins. Through its ability to integrate genetic data with mechanistic modeling, computational biophysics significantly accelerates the discovery of precision treatments for breast, lung, and other cancers.
Our research focuses on virtual screening of natural product compound libraries to identify potential inhibitors targeting proteins encoded by the top 20 mutated genes in breast and lung cancers. Natural products offer a diverse array of bioactive molecules with unique chemical structures, making them invaluable in drug discovery. By docking these compounds onto selected proteins associated with critical mutations, we aim to uncover new inhibitors that can modulate aberrant protein functions, potentially leading to effective therapies for these prevalent cancers.
Molecular Docking of Urease Inhibitors: A Step Towards New Treatments for Gastric Ulcers
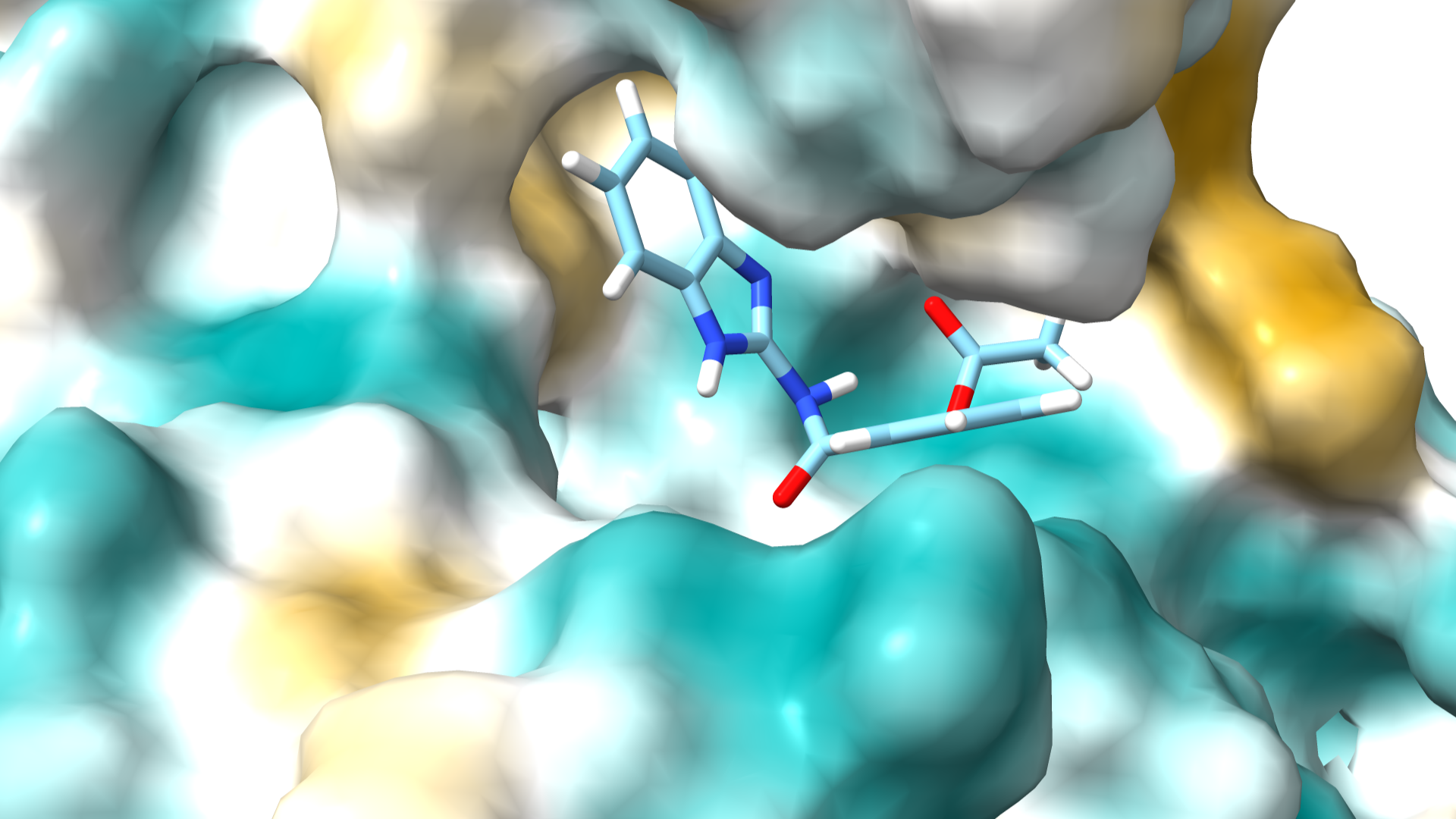
Urease is a metalloenzyme that catalyzes the hydrolysis of urea into ammonia and carbon dioxide, playing a vital role in nitrogen metabolism. Produced by pathogenic bacteria like Helicobacter pylori, urease raises local pH levels, facilitating bacterial survival in the acidic stomach. Understanding its structure and function is crucial for developing targeted inhibitors to combat its pathogenic effects.
Inhibiting urease offers significant therapeutic benefits, particularly for gastrointestinal health. Reduced urease activity lowers ammonia production, decreasing local pH and enhancing the efficacy of antibiotics against Helicobacter pylori. This leads to less gastric inflammation and ulcer formation. Additionally, urease inhibitors can improve nitrogen use in agriculture, reducing environmental impact. Thus, effective urease inhibitors are promising for both medical and agricultural applications.
Our research group is conducting a study using AutoDock Vina to identify potential inhibitors from a local library of compounds. This in silico approach predicts binding affinities and interactions between compounds and urease, guiding the development of novel therapeutics aimed at improving treatment outcomes for gastric ulcers and related disorders.
Targeting Aldehyde Dehydrogenase: A Virtual Screening Approach Using Natural Product Databases
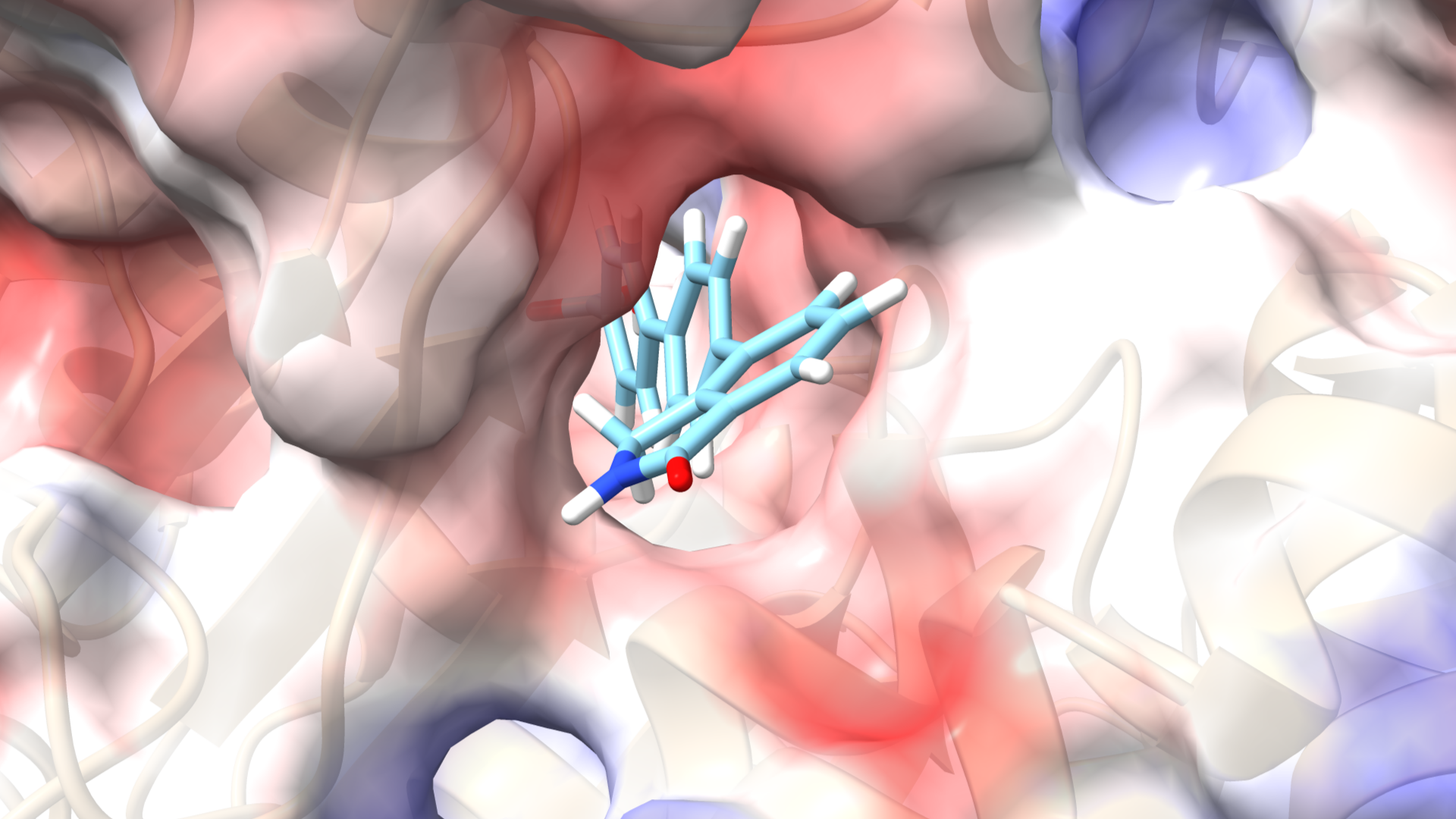
Aldehyde dehydrogenases (ALDHs) are a family of enzymes that catalyze the oxidation of aldehydes to carboxylic acids, playing a vital role in detoxification and metabolism. Found in various tissues, ALDHs help process toxic byproducts, including those from alcohol metabolism. Alterations in specific ALDH isoforms are linked to diseases such as cancer, neurodegenerative disorders, and metabolic syndromes.
Inhibiting ALDH proteins offers significant therapeutic benefits, particularly in cancer and neuroprotection. Some ALDH isoforms are linked to cancer stem cells, contributing to tumor growth and treatment resistance. Targeting these enzymes can enhance cancer therapy efficacy. Additionally, inhibiting specific ALDHs may reduce oxidative stress, making it a potential strategy for neurodegenerative diseases. Overall, ALDH inhibition can provide insights into metabolic pathways and address conditions involving aldehyde accumulation.
Our research group is conducting a virtual screening study using natural product databases to identify potential ALDH inhibitors. Utilizing AutoDock Vina, we will simulate docking interactions between these natural compounds and various ALDH isoforms to predict binding affinities. This in silico approach aims to discover novel inhibitors for therapeutic applications in cancer treatment and other ALDH-related conditions, leveraging the diversity of natural products.
